Department of Applied Chemistry, Graduate School of Enginnering, Osaka University* Unit of Biochemistry, Department of Medical Science, Faculty of Science, Rangsit University, Thailand** Department of Applied Chemistry, Graduate School of Engineering, Osaka University, Japan*** Department of Electrical Engineering, Osaka University, Japan**** Department of Material and Life Science, Osaka University, Japan***** Department of Cell Membrane Biology, Institute of Scientific and Industrial Research, Osaka University, Japan****** Department of Biochemistry, Faculty of Medicine, Chulalongkorn University, Thailand******* Department of Molecular Protozoology, Research Institute for Microbial Diseases, Osaka University, Japan********
○Keiji Tokuoka* Sudaratana R Krungkrai** Yukiko Kusakari*** Tsuyoshi Inoue*** Hiroaki Adachi**** Hiroyoshi Matsumura*** Kazufumi Takano***** Satoshi Murakami****** Yusuke Mori**** Yasushi Kai*** Krungkrai Jerapan******* Toshihiro Horii********
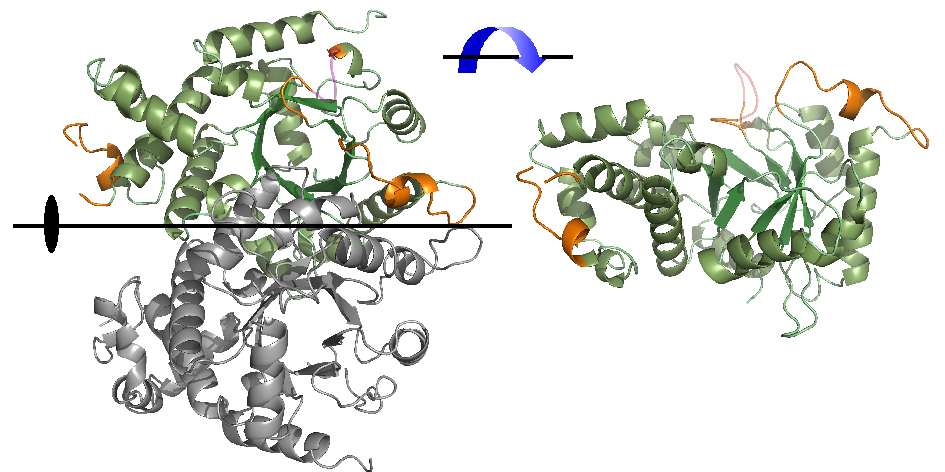
Orotidine 5'-monophosphare (OMP) dacarboxylase catalyze the decarboxylation reaction to produce uridine 5'-monophosphate (UMP) in the final step of UMP synthesis. The crystal structure of recombinant malarian Plasmodium falciparum OMP decarboxylase has been determined in the apo form at 2.6 Å resolution. OMP decarboxylase is a dimmer of two identical subunits. Each monomer consists of a triosephosphate isomerase barrel and contains an active site that is located across one end of the barrel and near the dimmer interface. Comparing with OMP decarboxylase from P.Vivax in complex with substrate, we found the remarkable structural change of loop region (#264-277) that locates near the active site. Plasmodium falciparum is the causative agent of the most lethal and sever form of human malaria. Chemotherapy of malaria is available, but is complicated by both adverse effects and widespread resistance to most of the currently available anti-malaria drugs. The malaria parasite depends on de novo synthesis of pyrimidine nucleotides, whereas the human host has the ability to synthesize them by both de novo and salvage pathways. Anti-malaria drugs design by using the structure of OMP dacarboxylase is in progress.
